Data Engineering: Crop big raster files with Python and rasterio
Prototyping of EO data processing and analysis scripts could be painful due to the size of images.
As a remote sensing specialist and a big fan of MacBook. I usually work in a coffe-house with my laptop and I must find a way to process data with limited RAM access. My laptop has 8 GB of memory. Typical Earth Observation product (image, classification map etc.) has a size of 1 GB. Someone could say that 8 GB is more than enough to read this file but processing it is a different piece of cake. It is possible to work with a big rasters when specialized software is used (QGIS). But when I’m working in Jupyter Notebook and I’m using numpy processing algorithms then RAM demand is close to my MacBook limit or exceeds it.
Fortunately there are two methods to overcome this problem. The first method is to read only part of an image.
The second method is to crop big image into smaller parts and store them for further processing. I’ve described it in this article.
Image cropping is very important for reproducibility of the processing piepline where, theoretically, we may validate every processing step. The major drawback of this method is a problem with a data storage. It is an exchange between RAM and the hard drive memory. MacBook’s hard drive memory volume is very small so I must use external drive to store remote sensed images but at the same time.
Cropping of georeferenced image may be performed with rasterio library. It is the must have Python tool of a remote sensing specialist. When you prototype machine learning systems where satellite imagery is analyzed then rasterio is a better choice than GDAL. (Ok, in practice it depends, some data formats are not well suited for rasterio… or rasterio is not well suited for them). We need two more libraries for our purpose: numpy (linear algebra and matrix operations) and matplotlib (data visualization).
Custom Python class to perform a clipping operation is presented in the listing below. Link to full implementation with docstrings is given at the end of article.
import numpy as np
import rasterio as rio
import affine
class ImageForClipModel:
# Get band, band shape, dtype, crs and transform values
def __init__(self, image_address):
with rio.open(image_address, 'r') as f:
self.band = f.read(1)
self.crs = f.crs
self.b_trans = f.transform
self.band_shape = self.band.shape
self.band_dtype = self.band.dtype
self.clipped_images = []
self.clipped_addresses = []
# Function for clipping band
def clip_raster(self, height, width, buffer=0,
save_mode=False, prefix='clipped_band_',
pass_empty=False):
r_pos = 0
while r_pos < self.band_shape[0]:
col_pos = 0
while col_pos < self.band_shape[1]:
clipped_image = self.band[r_pos:r_pos+\
height,
col_pos:col_pos + width]
# Check if frame is empty
if pass_empty:
if np.mean(clipped_image) == 0:
print('Empty frame, not saved')
break
# Positioning
tcol, trow = self.b_trans * \
(col_pos, r_pos)
new_transform = affine.Affine(self.b_trans[0],
self.b_trans[1],
tcol,
self.b_trans[3],
self.b_trans[4],
trow)
image = [clipped_image, self.crs, new_transform,
clipped_image.shape[0],
clipped_image.shape[1],
self.band_dtype]
# Save or append into a set
if save_mode:
filename = prefix + 'x_' + \
str(col_pos) + '_y_' + \
str(r_pos) + '.tif'
with rio.open(filename, 'w',
driver='GTiff',
height=image[3], width=image[4],
count=1, dtype=image[5],
crs=image[1],
transform=image[2]) as dst:
dst.write(image[0], 1)
self.clipped_addresses.append(filename)
else:
self.clipped_images.append(clipped_image)
# Update column position
col_pos = col_pos + width - buffer
# Update row position
r_pos = r_pos + height - buffer
if save_mode:
print('Tiles saved successfully')
return self.clipped_addresses
else:
print('Tiles prepared successfully')
return self.clipped_images
With this class it is possible to divide image into georeferenced quadrants. Example of the code usage (from Jupyter Notebook):
%matplotlib inline
import matplotlib.pyplot as plt
test_image = /
'raster_clip/LC08_L1TP_190022_20180413_20180417_01_T1_sr_band5.tif'
# Show test image
with rio.open(test_image, 'r') as src:
img = src.read(1)
img = img.astype(np.float32)
img = img / np.max(img)
img[img <= 0] = np.nan
plt.figure(figsize=(12, 12))
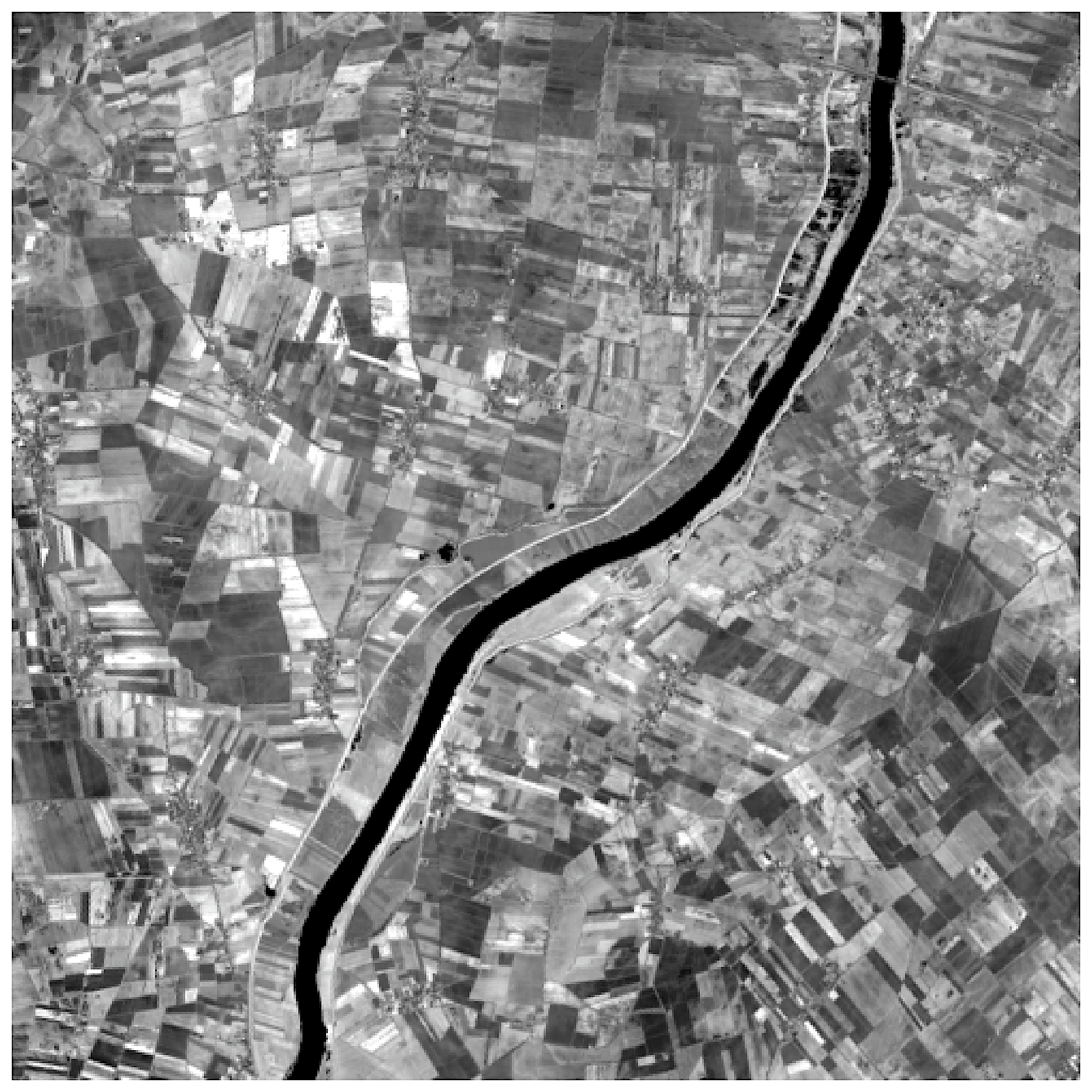
plt.imshow(img, cmap='gray')
plt.show()
>> Output:

clipper = ImageForClipModel(image_address=test_image)
clipper.clip_raster(height=500,
width=500,
buffer=0,
save_mode=True,
prefix='raster_clip/clipped_band_',
pass_empty=False)
>> Output:
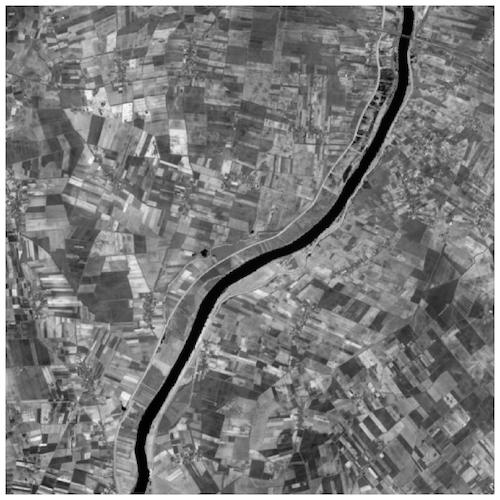

References
Script: https://github.com/szymon-datalions/geoprocessing/blob/master/rasters/clip_into_quadrants.py